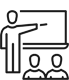
浙江大学与招商局集团在香港签署战略合作协议
4月23日下午,浙江大学与招商局集团在香港签署战略合作协议,名校名企合作迈上新台阶。 浙江大学党委书记任少波、招商局集团董事长缪建民出席并见证签约。浙江大学党委常委、副校长…
- 2024-04-23 【中新网】中国历代绘画大系亮相意大利威尼斯双年展
- 2024-04-23 浙江大学校长杜江峰院士率代表团访问意大利
- 2024-04-23 国际公共卫生论坛暨WHO胜任力导向公共卫生教育国际会议...
- 2024-04-23 浙江大学科技期刊主编联席会议在紫金港校区举行
- 2024-04-22 第三届全国前沿交叉研究院院长联席会年会在浙大举行
国家开放大学来浙江大学继续教育学院调研交流
4月19日,国家开放大学党委书记、校长王启明,副校长范贤睿一行来浙大干训基地、继续教育学院调研。学院纪委书记、副院长沈燎接待。双方相关部门负责人出席会议。 王启明表示,国家开放大学与…
- 2024-04-18 浙江大学继续教育学院现场教学基地(泾县)揭牌仪式在宣城市举行
- 2024-04-18 杭州市人大常委会副秘书长魏虹一行来澳门人巴黎人1797调研交流
- 2024-04-12 澳门人巴黎人1797获得浙江大学2023年度中层领导班子考核优秀
- 2024-04-11 【人民日报】#浙大紫藤花长廊浪漫氛围拉满# 华家池校区的紫藤...
- 2024-04-09 浙江大学继续教育学院发展联络顾问聘任仪式在华家池校区举行
2024-04-23 干部履职 | 特检院领导干部综合能力提升培训专题
2024-04-22 干部履职 | 外事侨务干部业务能力提升培训专题
2024-04-09 干部履职 | 统战系统干部政产学研用创新培训专题
2024-04-09 干部履职 | 气象局干部综合能力提升专题培训专题
2024-04-09 干部履职 | 无线电管理局干部素质能力提升培训专题
2024-03-22 干部履职 | 机构编制系统领导干部能力提升培训专题
2024-03-22 干部履职 | 烟草专卖系统干部综合素质提升培训专题
-
潍坊市在浙大举办科技创新赋能现...
2024-04-25
-
张家湾镇人民政府在浙大开展乡村...
2024-04-25
-
平顶山高新区科技创新局在浙大举...
2024-04-25
-
哈尔滨市工商联在浙大举办青年企...
2024-04-25
-
商洛市商州区委组织部在浙大举办...
2024-04-25
-
德阳市委组织部、市委教育工委在...
2024-04-25
-
察布查尔县委组织部在浙大举办铸...
2024-04-25
-
安顺市委组织部委托浙大举办新型...
2024-04-22
-
莆田市科技局委托浙大举办科技创...
2024-04-22
-
石家庄市直机关党组织书记赴浙大...
2024-04-22
2024-04-24 企业培训 | 企业集团优秀班组长能力提升专题培训班
2024-04-03 企业培训 | 银行柜员柜台操作技能提升与风险防范培训班
2024-04-03 企业培训 | 银行客户经理综合素质提升专题培训班
2024-04-18 企业培训 | 保险公司营销骨干成长计划培训班
2024-04-17 企业培训 | 投资集团公司中高层管理人员能力提升培训班
2024-04-17 企业培训 | 国有企业董事、监事、高级管理人员能力提升专题培...
2024-04-17 企业培训 | 国有企业人力资源管理专题培训班
-
山东省国有资产投资控股有限公司委托浙...
2024-04-24
-
龙城发展投资集团有限公司管理人员赴浙...
2024-04-11
-
保利集团“青马工程”第三期结业仪式暨...
2024-04-10
-
中国东方航空集团委托浙大举办数字化转...
2024-04-03
-
新疆银行与浙江大学共办运营业务能力提...
2024-03-28
-
中国工商银行舟山分行中青年干部、优秀...
2023-12-12
-
国航股份飞行总队工会干部走进浙江大学...
2023-12-05
-
中国铁建大桥工程局集团有限公司委托浙...
2023-11-30
-
国航股份浙江分公司、上海分公司委托浙...
2023-11-28
-
广晟控股集团委托浙大举办红色主题教育...
2023-11-27
开班动态
-
中国地质大学(北京)委托浙大举办处级...
2024-04-19
-
空军军医大学委托浙大举办研究生导师能...
2024-04-03
-
华中师范大学委托浙大举办“华大先锋·...
2024-04-03
-
四川传媒学院骨干教师及专业带头人赴浙...
2024-03-04
-
哈尔滨工业大学继续教育学院干部赴浙江...
2024-03-15
-
西安理工大学骨干教师走进浙江大学培训...
2024-01-19
-
四川音乐学院委托浙大举办高层次人才强...
2024-01-15
-
华南农业大学纪检监察干部走进浙江大学...
2023-11-29
更多>现场教学
嘉兴南湖红色教育基地
全国爱国主义教育示范基地
浙东(四明山)抗日根据...
浙东(四明山)抗日根据地...
 办公网
办公网 0571-86971085
0571-86971085






















 2024-04-20
2024-04-20

























































 2024-04-25
2024-04-25















 校园映像
校园映像